Автоматизована детекція та класифікація мі-кроскопічних фотозображень діатомових во-доростей
Основний зміст сторінки статті
Анотація
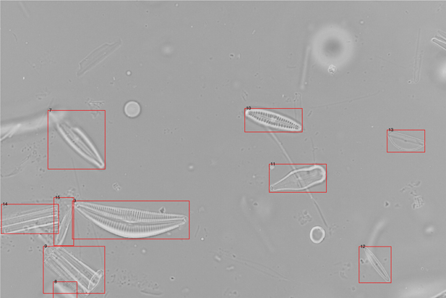
Досліджені алгоритми автоматизованої детекції та визначення видів діатомових водоростей на мікроскопічних фотозображеннях для завдань екологічного моніторингу. Запропоновано метод порогової обробки фотозображень для ефективної детекції зображень із різною контрастністю. Навчена та досліджена згорткова нейронна мережа для визначення видів діатомових водоростей на фотозображеннях.
Блок інформації про статтю

Ця робота ліцензується відповідно до Creative Commons Attribution 4.0 International License.
Автори, які публікуються у цьому журналі, погоджуються з наступними умовами:
- Автори залишають за собою право на авторство своєї роботи та передають журналу право першої публікації цієї роботи на умовах ліцензії Creative Commons Attribution License, котра дозволяє іншим особам вільно розповсюджувати опубліковану роботу з обов'язковим посиланням на авторів оригінальної роботи та першу публікацію роботи у цьому журналі.
- Автори мають право укладати самостійні додаткові угоди щодо неексклюзивного розповсюдження роботи у тому вигляді, в якому вона була опублікована цим журналом (наприклад, розміщувати роботу в електронному сховищі установи або публікувати у складі монографії), за умови збереження посилання на першу публікацію роботи у цьому журналі.
- Політика журналу дозволяє і заохочує розміщення авторами в мережі Інтернет (наприклад, у сховищах установ або на особистих веб-сайтах) рукопису роботи, як до подання цього рукопису до редакції, так і під час його редакційного опрацювання, оскільки це сприяє виникненню продуктивної наукової дискусії та позитивно позначається на оперативності та динаміці цитування опублікованої роботи (див. The Effect of Open Access).
Посилання
Bukhtiyarova L. N. “Bacillariophyta v biomonitoringe rechnykh ekosistem. Covremennoye sostoyaniye i perspektivy ispol'-zovaniya [Bacillariophyta in the biomonitoring of river ecosystems. Current state and prospects of use]”, Al'gologiya, vol. 9, no. 3, pp. 89–103, 1999.
Bukhtiyarova L. N. “Classification of diatom algocoenoses as a useful tool in river biomonitoring. In: J. Prygiel, B. A. Whitton, J. Bukowska (eds) Use of algae for Monitoring Rivers. III”. – Agence de l'Eau Artois-Picardie, p. 114–121, 1999. URL: https://www.researchgate.net/publication/282974818_Classification_of_diatom_algocoenoses_as_a_useful_tool_in_river_biomonitoring
Treguer R., Nelson D. M., Van Bennkom A. J. “The silica balance in the world ocean: a reestimate”, Science, vol. 268, p. 375–379, 1995. URL: https://www.researchgate.net/ publication/6092964_The_Silica_Balance_in_the_World_Ocean_A_Reestimate
The European Parliament and the Council of the European Union. Directive 2000/60/EC. Establishing a Framework for Community Action in the Field of Water Policy; Official Journal of the European Community: Maastricht, the Netherlands, Work Safety Introduction, 2000. URL: https://injuryfacts.nsc.org/work/work-overview/work-safety-introduction/.
Lobo E., Heinrich C., Schuch M., Wetzel C., Ector L. “Diatoms as Bioindicators in Rivers”, River Algae, vol. 06, p. 245–271, 2016. https://doi.org/10.1007/978-3-319-31984-1_11
Wang Yi-K., Stevenson R., Sweets R., DiFranco J. “Developing and testing diatom indicators for wetlands in the Casco Bay watershed, Maine, USA” Hydrobiologia, vol. 561, no. 191, p. 245–271. 2006. DOI: 10.1007/s10750-005-1614-2
Stevenson R., Pan Y., Van Dam H. “Assessing environmental conditions in rivers and streams with diatoms”. In: Smol J. P. and Stoermer E. F. (Eds.) – The Diatoms: Applications for the Environmental and Earth Sciences. Cambridge, UK, Cambridge University Press, p. 57–85, 2010. DOI: 10.1017/CBO9780511763175.005.
Mayama S., Katoh K., Omori H., Seino S., Osaki H., Julius M., Lee J. H., Cheong C., Lobo E. A., Witkowski A., Srivibool R., Muangphra P., Jahn R., Kulikovskiy M., Hamilton P. B., Ya-Hui G., Ector L., Soeprobowat T. R. “Progress toward the Construction of an International Web-based Educational System Featuring an Improved "SimRiver" for the Understanding of River Environments”, Asian Journal of Biology Education, vol. 5, p. 2–14. 2011. URL: http://www.u-gakugei.ac.jp/~mayama/PDF/Mayama et al_2011_SimRiver_AJBE.pdf
Dimitrovski I., Kocev D., Loskovska S., Dzeroski S., “Hierarchical classification of diatom images using ensembles of predictive clustering trees”, Ecological Informatics, vol. 7, p. 19–29, 2012. URL: http://kt.ijs.si/DragiKocev/ECOINF/ECOINF_diatom_images.pdf
Bueno G., Deniz O., Pedraza A., Ruiz-Santaquiteria J., Salido J., Cristóbal G., Borrego-Ramos M., Blanco S., “Automated Diatom Classification (Part A): Handcrafted Feature Approaches,” Applied Sciences, vol. 7, no. 8, p. 753, 2017. URL: http://digital.csic.es/bitstream/10261/153574/1/applsci-07-00753.pdf
Pedraza A., Bueno G., Deniz O., Cristóbal G., Blanco S., Borrego-Ramos M. Automated Diatom Classification (Part B): A Deep Learning Approach, – Applied Sciences vol 7, p. 460, 2017. DOI: 10.3390/app7050460
Quantitative image analysis of microstructures. Exner H. E., Hougardy H. P. (Eds). DGM Informations‐gesellschaft Verlag, Oberursel, 235 pp., 1988. ISBN 3‐88355‐132‐5 DOI: 10.1002/adma.19900020216
Glasbey C. A., Horgan G. W. “Image analysis for the biological sciences”, Wiley J., Sons, Inc. New York, NY, USA, 230 pp. 1995. ISBN: 0-471-93726-6 DOI: 10.1017/S0021859600088924
Zheng J., Zhang D., Huang K., Sun Y., Tang S., “Adaptive windowed range-constrained Otsu method using local information”, Journal of Electronic Imaging, vol. 25, p. 013034 , 2016, DOI: 10.1117/1.JEI.25.1.013034
Jung A. “Imgaug”, 2017. URL: https://github.com/aleju/imgaug.
Iandola F. N., Han S., Moskewicz M. W., Ashraf K., Dally W. J., Keutzer K. “SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and
Simonyan K., Zisserman A. “Very deep convolutional networks for large-scale image recognition”. arXiv preprint, 2014. arXiv: 1409.1556